The disease tuberculosis (TB) which is caused by the bacterium Mycobacterium Tuberculosis, is an infectious disease listed among the top 10 causes of fatality worldwide. It killed up to 1.4 million people in 2019 (WHO), thus making it as one of the top 10 causes of death.
The enoyl-acyl carrier protein reductase (InhA) from M. Tuberculosis, is one of the key enzymes which is responsible for synthesis of essential mycolic acids in the mycobacterial cell wall. Therefore, InhA is the key target for anti-tubercular drug. Multi drug resistant TB remains a public health crisis, thereafter, the studies of potent and novel inhibitors are of great interest.
Therefore, Assoc. Prof. Dr. Pornpan Pungpo (Department of Chemistry, Faculty of Science, Ubon Ratchathani University) has been interested in the development of the potent and novel inhibitors.
3NP (Figure 1A) has been extracted from endophytic fungi, Phomopsis sp. usia5 from plant hosts located in Thailand. It is a bioactive compound and proposed as an antitubercular agent. It is a promising antimycobacterial agent, thus exhibiting minimum inhibitory concentration (MIC) values of 50.0 µg/mL and 12.5 µg/mL against M. tuberculosis H37Ra and M. tuberculosis H37Rv, respectively. 3NP showed half maximal inhibitory concentration (IC50) values for InhA inhibition of 71.2 μM, compared to a value for InhA inhibitor triclosan of 5.32 μM.
Macromolecular Crystallography (MX) has been carried out for studying the three dimensional structure of holo structure of InhA from M. Tuberculosis (Figure 1B). Dr. Chomphunuch Songsiriritthigul (Synchrotron Light Research Institute, SLRI) performed crystal structure analysis under the supervision of Prof. Dr. James Spencer (School of Cellular and Molecular Medicine, University of Bristol, United Kingdom).
(A) (B)
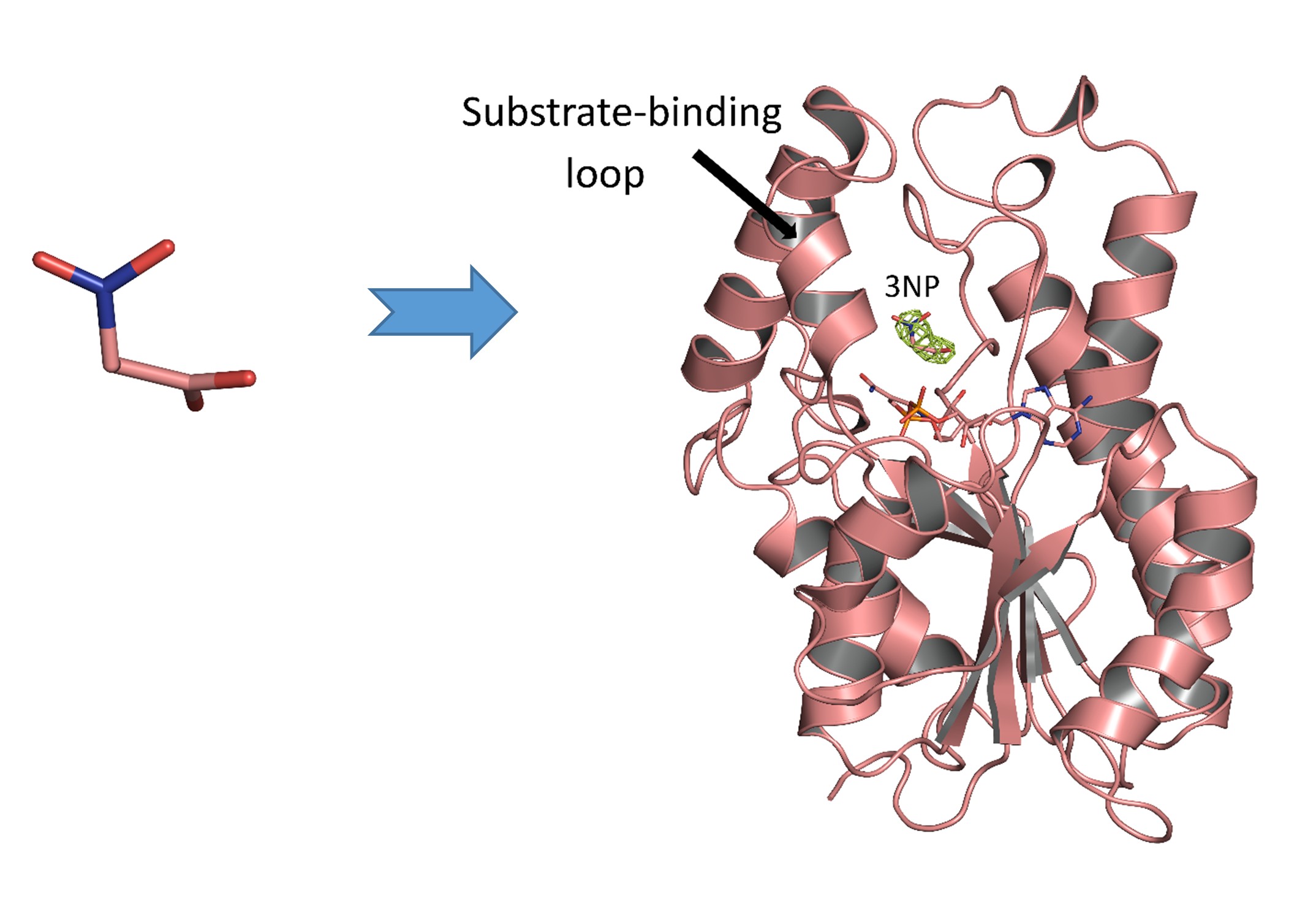
Figure 1 (A) Chemical structure of 3-nitropropanoic acid (3NP), a bioactive compound. (B) The overall structure of InhA-NAD+-3NP complex (PDB entry 7E48).
Citation: PROTEINS: Structure, Function, and Bioinformatics. 2022; 90(3): 898-904. DOI: 10.1002/prot.26268.
Crystal structure of the complex contains the InhA substrate-binding loop in an ordered, open conformation with Tyr158, a catalytically important residue whose orientation defines different InhA substrate/inhibitor complexes, in the “out” position (Figure 2A). 3NP occupies a hydrophobic binding site adjacent to the NAD+ cofactor and close to that utilized by the diphenyl ether triclosan (Figure 2B), but binds predominantly via electrostatic and water-mediated hydrogen-bonding interaction with the protein backbone and NAD+ cofactor.
(A) (B)
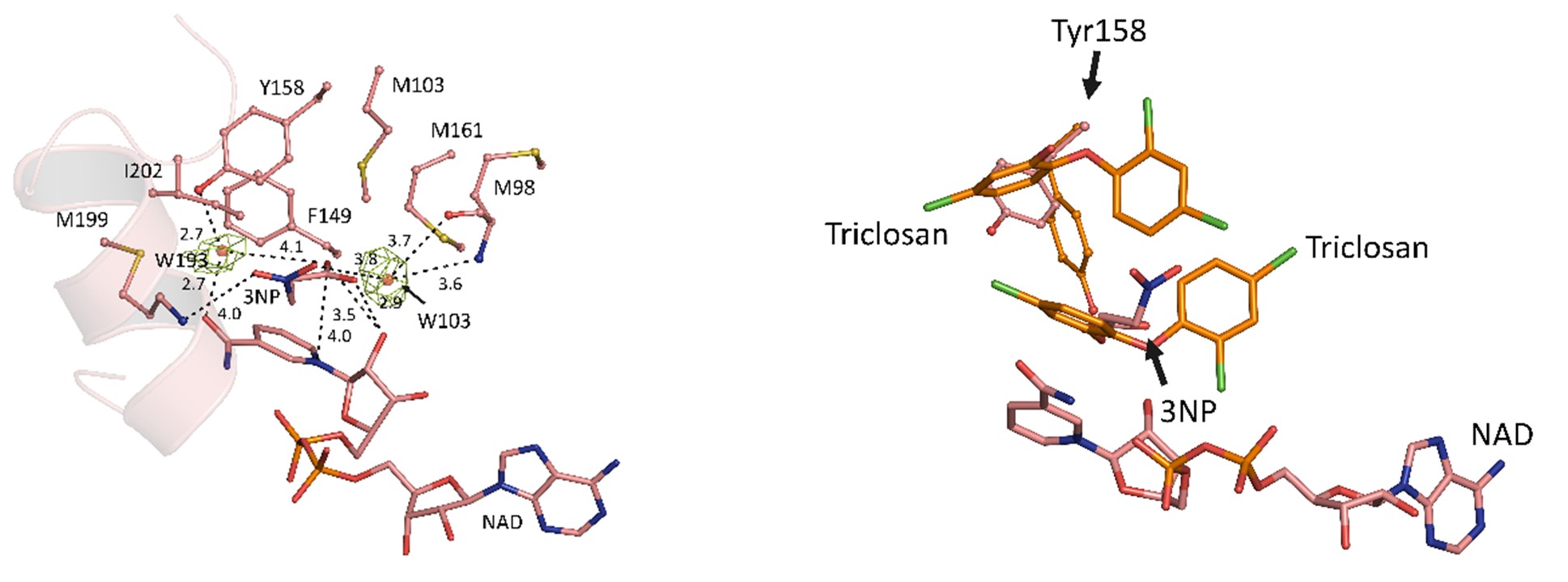
Figure 2 (A) The structural components of InhA-NAD+-3NP complex (PDB entry 7E48, salmon). Water-mediated hydrogen bonds which stabilized both 3NP and NAD+ are shown. (B) The superposed structures of InhA-NAD+-3NP (PDB entry 7E48, salmon) and InhA-NAD+-triclosan (PDB entry 1P45, orange).
Citation: PROTEINS: Structure, Function, and Bioinformatics. 2022; 90(3): 898-904. DOI: 10.1002/prot.26268.
Therefore, 3NP retains value as a starting point as a pharmacophore for potential antituberculosis drugs owing to its ready availability from natural and commercial sources, the potential for developing derivatives that may reduce toxicity and improve InhA inhibitory activity.
The lack of involvement of specific amino acid side chains in 3NP binding suggests that compounds exploiting these interactions might show reduced susceptibility to emergence of resistance by mutation of the InhA active site. These data indicate that further exploration and compound optimization, for example derivatization of 3NP, may be warranted to exploit these findings in potential antituberculosis agents.
----------------------------------------------------------------------------------------------
Written by Chomphunuch Songsiriritthigul (Beamline Scientist, SLRI)
References
- Chomcheon, P, Wiyakrutta, S, Sriubolmas, N, Ngamrojanavanich, N, Isarangkul, D, Kittakoop, P. 3-Nitropropionic Acid (3-NPA), a Potent Antimycobacterial Agent from Endophytic Fungi: Is 3-NPA in Some Plants Produced by Endophytes?. J Nat Prod. 2005; 68: 1103-1105.
- Ganihigama, DU, Sureram, S, Sangher, S, Hongmanee, P, Aree, T, Mahidol, C, Ruchirawat, S, Kittakoop, P. Antimycobacterial activity of natural products and synthetic agents: Pyrrolodiquinolines and vermelhotin as anti-tubercular leads against clinical multidrug resistant isolates of Mycobacterium tuberculosis. Eur J Med Chem. 2015; 89: 1-12.
- Kamsri, P, Hanwarinroj, C, Naruedon, P, et al. Discovery of New and Potent InhA Inhibitors as Antituberculosis Agents: Structure-Based Virtual Screening Validated by Biological Assays and X‑ray Crystallography. J Chem Inf Model. 2019; 60: 226-234.
- Songsiriritthigul, C., Hanwarinroj, C., Pakamwong, B., Srimanote, P., Suttipanta, N., Sureram, S., Suttisintong, K., Kamsri, P., Punkvang, A., Spencer, J., Kittakoop, P., Pungpo, P. Inhibition of Mycobacterium tuberculosis InhA by 3-nitropropanoic acid. Proteins: Struct Funct Bioinf. 2022; 90(3): 898-904. DOI: 10.1002/prot.26268.
- World Health Organization. https://www.who.int/news-room/fact-sheets/detail/tuberculosis. Accessed: February 2021.

